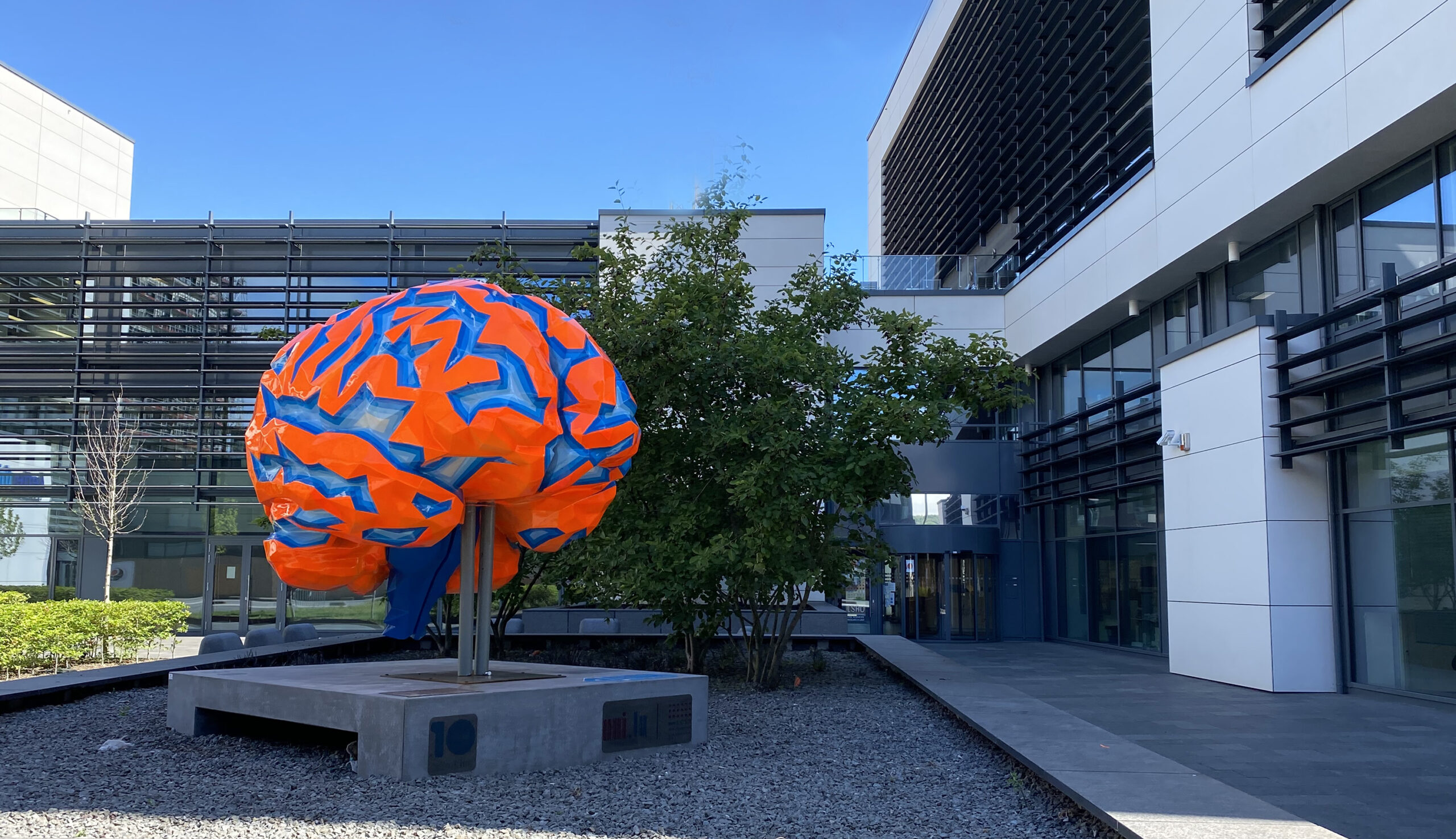
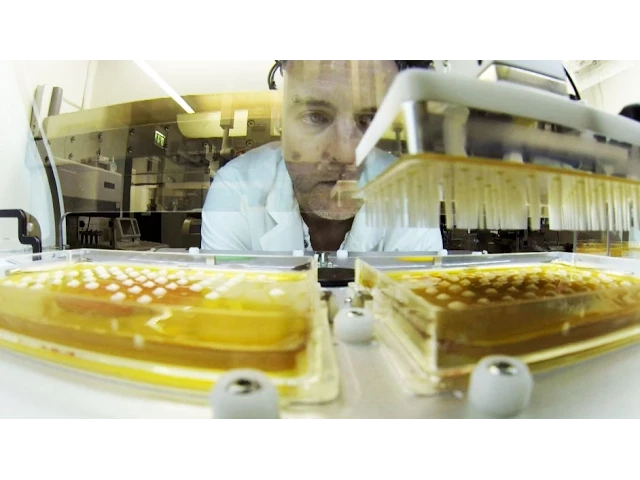
A glimpse into our research
Our research focuses on neurodegenerative diseases such as Alzheimer’s or Parkinson’s. When do they start to develop? What is the role of genetics? How are they influenced by lifestyle, diet or environment? How do all these factors interact? How can we slow down the progression of neurodegenerative diseases or even prevent them? We are tirelessly searching for answers, for the benefit of patients.
LCSB numbers:
-
270Staff members
-
18Research groups
Our research areas
News
-
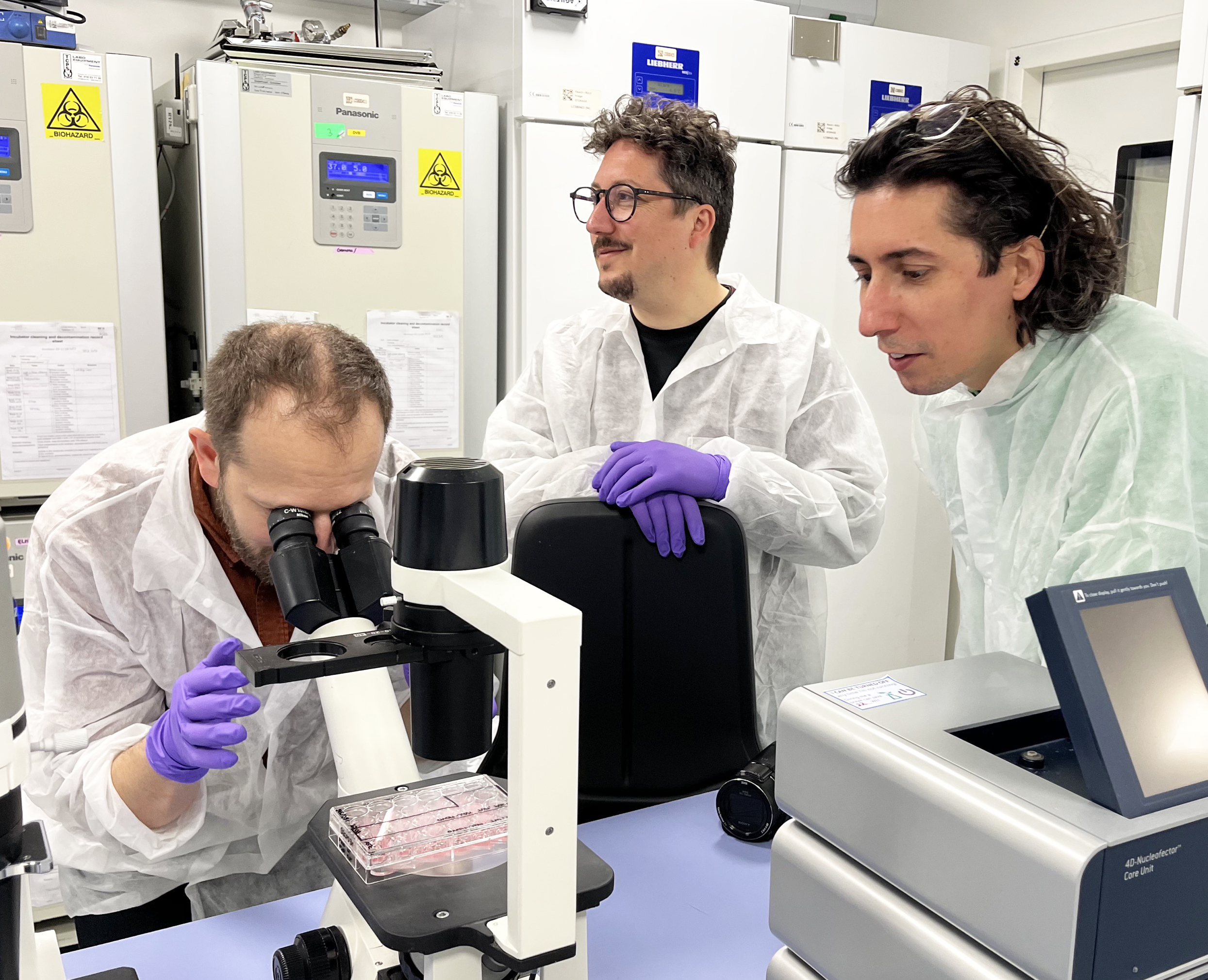

Art meets science: A first artist residency at the university
Campus Life, OutreachLife Sciences & MedicineLearn more -

Rare disease research and outreach at the LCSB – Let’s care for rare!
Outreach, ResearchLife Sciences & MedicineLearn more
Events
20 years dedicated to students, research and society
2023 marks the 20th anniversary of the University of Luxembourg, and we have many reasons to celebrate. What began as an academic startup, is now an international research university with a distinctly multilingual and interdisciplinary character. Our alumni community has grown to more than 14,000 graduates with many of them staying in Luxembourg after securing jobs or embarking on their own entrepreneurial journeys, making the country even more competitive. Others have carved out impressive careers abroad.
In this playlist, we’ll release weekly videos in which you’ll meet a total of 20 graduates from the past 20 years and learn how their time at the University helped them get where they are today.